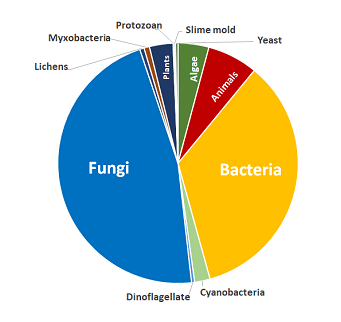
概述
用于天然化合物研究中的化合物筛选和其他应用的天然产物数据库
著名的 AntiBase Library 数据库最初由 Hartmut Laatsch 博士开发,是一个包含 105,300 多种化合物的综合性天然产物数据库。随着最新版本的发布,我们又增列了 9,500 多种化合物!
该数据库在科学文献中被广泛引用,规模持续扩展,是代谢组学与药物研发领域的高效筛选工具——其收录的天然产物及衍生物可加速癌症、糖尿病、各类疾病以及抗菌/抗真菌疗法的研发进程。
使用内置的 Wiley ChemWindow 软件可以安全地搜索数据库集,更有多款强大的工具,助您建立自己的数据库、绘制结构图等。
覆盖全面。 |
数据经过精心整理。 |
数据挖掘工具功能强大。 |
|
凭借更广泛的化合物覆盖范围,加速各类天然产物的研究进程—这一丰富资源将助力您的科研发现。 |
在 Wiley,我们将数据质量放在首位,为您提供可信赖的结果,帮助您做出明智的决策。 |
功能强大的数据挖掘工具,助力获得更加深入的洞察,支持构建并整合自有数据及其他拓展功能! |
应用:
- 将其整合至制药和生物技术领域的高通量筛选(HTS)和去重复工作流程中,快速鉴定已研究过的化合物,从而优化具有抗菌或特定药效的新型化合物筛选流程。
- 可用于各行各业,如食品、化妆品、农业、杀虫剂以及天然产物普遍存在的其他应用,也可用于学术项目和研究。
2025 版有哪些新增功能
数据库集总收录量扩大了 10%,新增了 9,500 多种化合物,其中包括最新发布的“Natural Product Atlas”中的化合物。
来源可靠的靠谱数据
Wiley是科学数据的权威来源。我们著名的数据库根据严格的协议进行处理,确保其达到最优品质。这些鉴定程序从数据采集开始,贯穿整个数据库开发过程。我们从可信赖的合作伙伴处获取数据,且所有数据需经过全面审查后才能纳入我们的数据库。
主要功能
| 数据经过精心整理和验证 | 数据取自一次和二次文献,经过仔细检查和验证,数据字段中提供了主要参考文献和来源。 |
| 宝贵的元数据可帮助您缩小结果范围 |
除化学结构外,每条记录还包含了可搜索的元数据(如有),例如:
|
| 光谱数据为鉴定助力 |
|
| 强大的数据挖掘和数据库工具 |
内置 Wiley ChemWindow 软件,用于搜索数据库集,更有多款强大的工具,助您建立自己的数据库、绘制结构图等。 |
库技术指标
|
|
|
兼容性
- Wiley Identifier of Natural Products 内置并兼容 KnowItAll ChemWindow 版软件。
- 也可与您正在使用的其他版本的 KnowItAll 软件一起使用。
订购信息
|
Product Code |
Product |
|
978EALDB04188 |
Wiley 天然产物标识符 – AntiBase 库 + ChemWindow (Wiley Identifier of Natural Products (AntiBase Library + ChemWindow Software)) (按年订阅) |
|
978EALDB05529 |
Wiley 天然产物标识符 – AntiBase 库 + ChemWindow (Wiley Identifier of Natural Products (AntiBase Library + ChemWindow Software)) (按年续订) |
|
978EALDB04195 |
Wiley Identifier of Natural Products (AntiBase Library) (按年订阅)* |
|
978EALDB05512 |
Wiley Identifier of Natural Products (AntiBase Library) (按年续订)* |
*需要 KnowItAll 软件